Next generation sequencing continues to gain traction as a standard of care as it becomes less complex and less costly. Targeted sequencing, in which only a subset of genes are enriched and amplified for sequencing, is especially cost effective. Clinicians are using the technique to more quickly provide answers to oncologists to ensure that patients actually get the right therapy.
In addition to being less costly, targeted sequencing includes a number of other benefits. It requires a minimal amount of DNA input, and samples sequenced using this method can be turned around quickly. This method also enables sequencing regions of interest at higher coverage levels, thereby enabling identification of low-frequency variants, such as fusions.
To enable laboratories to take a targeted sequencing approach, Illumina provides the AmpliSeq for Illumina Focus assay. This assay accommodates DNA or RNA samples and investigates 52 genes with known relevance to solid tumors. It’s capable of detecting low Variant Allele Frequency somatic mutations and enables the study of multiple cancer types, including lung, colon, breast, ovarian, melanoma, and prostate. At PierianDx, we are proud to announce that we have expanded support of this assay and we have integrated it fully with our interpretation and reporting tool, Clinical Genomics Workspace. As a result, if you want to implement this assay, there is a standardized and rapid way for you to do so.
Overview of the Workflow
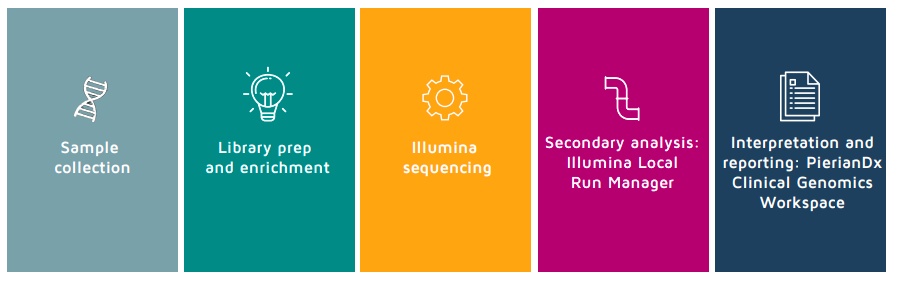
After samples are collected, library preparation is performed using a straightforward PCR-based protocol. The samples can then be sequenced using the Illumina MiniSeq™, MiSeq™, or iSeq™ systems, after which secondary analysis occurs on Illumina Local Run Manager. Users can then transfer .VCF files into PierianDx Clinical Genomics Workspace for tertiary analysis and production of a final clinical genomics report.
For more information, contact us.
References
1. Illumina. Benefits of NGS Targeted Resequencing. Accessed July 30, 2020.
2. AmpliSeq for Illumina Focus Panel | Combined DNA and RNA workflow. Illumina.com. https://www.illumina.com/products/by-type/sequencing-kits/library-prep-kits/ampliseq-focus-panel.html. Accessed July 30, 2020.

